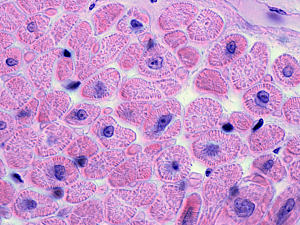
The early stages of allograft rejection after heart transplantation can be asymptomatic, so patients require post-transplant surveillance with endomyocardial biopsies (EMBs). There’s no standardized schedule, but most centers perform biopsies frequently for at least one to two years.
The gold standard for EMB evaluation is a manual histological examination of tissue slides. Interpretation is challenging because assessment is based on qualitative measurements, leading to subjective interpretations and consequently large inter-rater variability. In principle, different cardiac experts can come to a different diagnosis that affects the patients—causing either over- or under-treatment with immunosuppressive medications or unnecessary biopsy.
Researchers at Brigham and Women’s Hospital have created Cardiac Rejection Assessment Neural Estimator (CRANE), a form of artificial intelligence for automated screening of EMB. In Nature Medicine, they describe its performance and its potential to serve as an assistive diagnostic tool.
The authors of the report are Jana Lipkova, PhD, and Tiffany Y. Chen, MD, research fellows in the Department of Pathology, Faisal Mahmood, PhD, a researcher in the Division of Computational Pathology, and colleagues.
Description of CRANE
CRANE is a high-throughput, multitask, deep learning–enabled system that evaluates hematoxylin and eosin–stained whole-slide images. It simultaneously looks for acute cellular rejection, antibody-mediated rejection, concurrent cellular–antibody rejections, and Quilty B lesions. An additional network determines the grade (severity) of any rejection.
CRANE was developed on a dataset of 5,054 slides from 1,690 EMBs collected at the Brigham. The model is trained only with patient diagnosis as the only label.
Internal Test Cohort
In the hold-out test, CRANE performed very well on all tasks involved in detecting rejection. For overall performance, the area under the receiver operating characteristic curve (AUC) was 0.96 and accuracy was 90%.
Independent Test Cohorts
CRANE was tested on two independent cohorts from Turkey (1,717 slides from 585 patients) and Switzerland (123 slides from 123 patients). A variety of scanner vendors, biopsy protocols, and slide preparation procedures had been used.
Adapting the model from the Brigham cohort to these independent cohorts led to a drop in performance of 0.02 and 0.13 for AUC, and 2% to 14% for accuracy, similar to results for other deep learning models when applied to external datasets.
Comparison With Human Readers
The team performed two reader studies. The aim of the first one was to compare CRANE performance against the pathologists. The aim of the second one was to evaluate how useful CRANE can be for pathologists in their workflow.
Five board-certified pathologists from outside the Brigham were recruited to read 150 EMBs from the Turkish cohort, of which 50 had previously been determined by pathologists to be normal, and 100 showed rejection. To mimic CRANE’s process, the pathologists used hematoxylin and eosin–stained slides without immunohistochemistry analysis.
For all tasks, CRANE predictions were comparable to the human reads. The average agreement for rejection detection was κ=0.54 (moderate agreement) between individual pathologists and κ=0.64 (substantial agreement) between individual pathologists and CRANE.
Five different pathologists were then randomly assigned to one of two groups. One group assessed EMBs using only slides, while the second also used heatmaps that CRANE generates to predict the diagnostic relevance of each biopsy region on a slide. Four weeks later, the pathologists repeated the task using the opposite procedure.
The use of CRANE increased the accuracy for all tasks and reduced the assessment time for all readers.
Potential Applications of CRANE
Improved accuracy of rejection assessment with CRANE could reduce the number of unnecessary follow-up EMBs, an important outcome considering their expense and risks. Underestimation of rejection is also a problem and can result in treatment delays and poorer outcomes.
CRANE might also be used to screen for patients with advanced signs of rejections to allow prioritization of these patients and faster interventions (e.g., usually through medication to suppress the rejections).
Although CRANE’s performance in rejection grading is comparable to that of human experts, this task remains challenging, and the research team anticipates improvements to the model. Those might include integrating echocardiography results, cardiac hemodynamic measurements, and molecular biomarkers to improve risk stratification.